 PDF(1975 KB)
PDF(1975 KB)


Latent Space Embedding Methods for Chemical Molecules: Principles and Applications
Haotian Chen, Tao Yang, Xiaotong Liu
Prog Chem ›› 2025, Vol. 37 ›› Issue (10) : 1456-1478.
 PDF(1975 KB)
PDF(1975 KB)
 PDF(1975 KB)
PDF(1975 KB)
Latent Space Embedding Methods for Chemical Molecules: Principles and Applications
Effective representation of chemical molecules is the key to promoting chemical informatics and new material research and development. In recent years, data-driven molecular representation technology has been developed. Compared with traditional manually designed descriptors and graph structure analysis methods, it can effectively avoid noise and information redundancy, and provide support for efficient and accurate property prediction. Embedding representation has the characteristics of efficient information compression, data representation enhancement and semantic retention, and has been widely used in fields such as deep learning and data mining. Inspired by word embeddings in the field of natural language processing, researchers began to explore the application of similar methods to the construction of the latent space of chemical molecules, and proposed a variety of embedding methods for molecular property prediction and molecular structure generation. This review first elucidates the principles of general embedding technology in machine learning, and then sequentially discusses chemical element latent space representation methods and chemical molecule latent space embedding techniques. By examining the innovative applications of related technologies in natural language processing and graph embedding to molecular embeddings, the review reveals that current molecular embedding methods are gradually evolving towards multimodality, self-supervised learning, and dynamic modeling, and it outlines prospects for future research trends.
1 Introduction
2 Principles of embedding in machine learning
2.1 Word embedding
2.2 Graph embedding
2.3 Multimodal embedding
3 Element latent space representation methods
3.1 Attribute-based element representation
3.2 Element representation based on physicochemical knowledge
3.3 Data-driven element embedding
4 Advances in molecular latent space embedding
4.1 Traditional chemical feature-based molecular descriptors
4.2 Graph theory-driven molecular embedding
4.3 Data-driven molecular embedding
4.4 Multimodal molecular embedding
5 Conclusion and outlook
5.1 Current status and key technology
5.2 Future research prospects
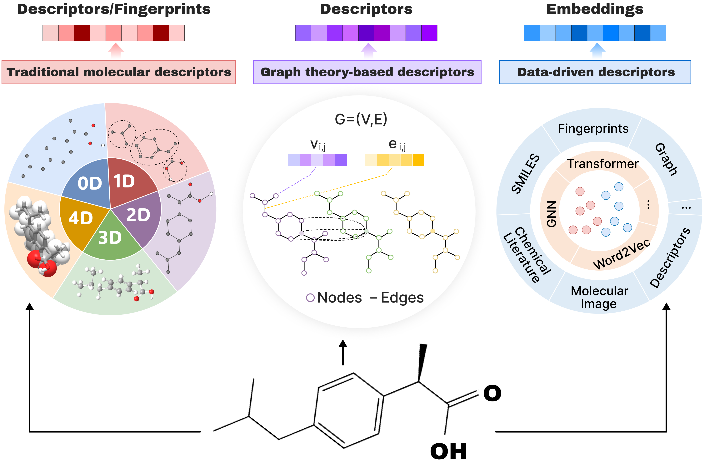
molecular embedding / machine learning / representation learning / property prediction / multimodality / self-supervised learning
| [1] |
|
| [2] |
|
| [3] |
|
| [4] |
|
| [5] |
|
| [6] |
|
| [7] |
|
| [8] |
|
| [9] |
|
| [10] |
|
| [11] |
|
| [12] |
|
| [13] |
|
| [14] |
|
| [15] |
|
| [16] |
|
| [17] |
|
| [18] |
|
| [19] |
|
| [20] |
|
| [21] |
|
| [22] |
|
| [23] |
|
| [24] |
|
| [25] |
|
| [26] |
|
| [27] |
|
| [28] |
|
| [29] |
|
| [30] |
|
| [31] |
|
| [32] |
|
| [33] |
|
| [34] |
|
| [35] |
|
| [36] |
|
| [37] |
|
| [38] |
|
| [39] |
|
| [40] |
|
| [41] |
|
| [42] |
|
| [43] |
|
| [44] |
|
| [45] |
|
| [46] |
|
| [47] |
|
| [48] |
|
| [49] |
|
| [50] |
|
| [51] |
|
| [52] |
|
| [53] |
|
| [54] |
|
| [55] |
|
| [56] |
|
| [57] |
|
| [58] |
|
| [59] |
|
| [60] |
|
| [61] |
|
| [62] |
|
| [63] |
|
| [64] |
|
| [65] |
|
| [66] |
|
| [67] |
|
| [68] |
|
| [69] |
|
| [70] |
|
| [71] |
|
| [72] |
|
| [73] |
|
| [74] |
|
| [75] |
|
| [76] |
|
| [77] |
|
| [78] |
|
| [79] |
|
| [80] |
|
| [81] |
|
| [82] |
|
| [83] |
|
| [84] |
|
| [85] |
|
| [86] |
|
| [87] |
|
| [88] |
|
| [89] |
|
| [90] |
|
| [91] |
|
| [92] |
|
| [93] |
|
| [94] |
|
| [95] |
|
| [96] |
|
| [97] |
|
| [98] |
|
| [99] |
|
| [100] |
|
| [101] |
|
| [102] |
|
| [103] |
|
| [104] |
|
| [105] |
|
| [106] |
|
| [107] |
|
| [108] |
|
| [109] |
|
| [110] |
|
| [111] |
|
| [112] |
|
| [113] |
|
| [114] |
|
| [115] |
|
| [116] |
|
| [117] |
|
| [118] |
|
| [119] |
|
| [120] |
|
| [121] |
|
| [122] |
|
| [123] |
|
| [124] |
|
| [125] |
|
| [126] |
|
| [127] |
|
| [128] |
|
| [129] |
|
| [130] |
|
| [131] |
|
| [132] |
|
| [133] |
|
| [134] |
|
| [135] |
|
| [136] |
|
| [137] |
|
| [138] |
|
| [139] |
|
| [140] |
|
| [141] |
|
| [142] |
|
| [143] |
|
| [144] |
|
| [145] |
|
| [146] |
|
| [147] |
|
| [148] |
|
| [149] |
|
| [150] |
|
| [151] |
|
| [152] |
|
| [153] |
|
| [154] |
|
| [155] |
|
| [156] |
|
| [157] |
|
| [158] |
|
| [159] |
|
| [160] |
|
| [161] |
|
| [162] |
|
| [163] |
|
| [164] |
|
| [165] |
|
| [166] |
|
| [167] |
|
| [168] |
|
| [169] |
|
| [170] |
|
| [171] |
|
| [172] |
|
| [173] |
|
| [174] |
|
| [175] |
|
| [176] |
|
| [177] |
|
| [178] |
|
| [179] |
|
| [180] |
|
| [181] |
|
| [182] |
|
| [183] |
|
| [184] |
|
| [185] |
|
| [186] |
|
| [187] |
|
| [188] |
|
| [189] |
|
| [190] |
EdwinChacko, RudraSondhi,
|
| [191] |
|
| [192] |
|
| [193] |
|
| [194] |
|
| [195] |
|
/
| 〈 |
|
〉 |